Actualités
17 avril 2024
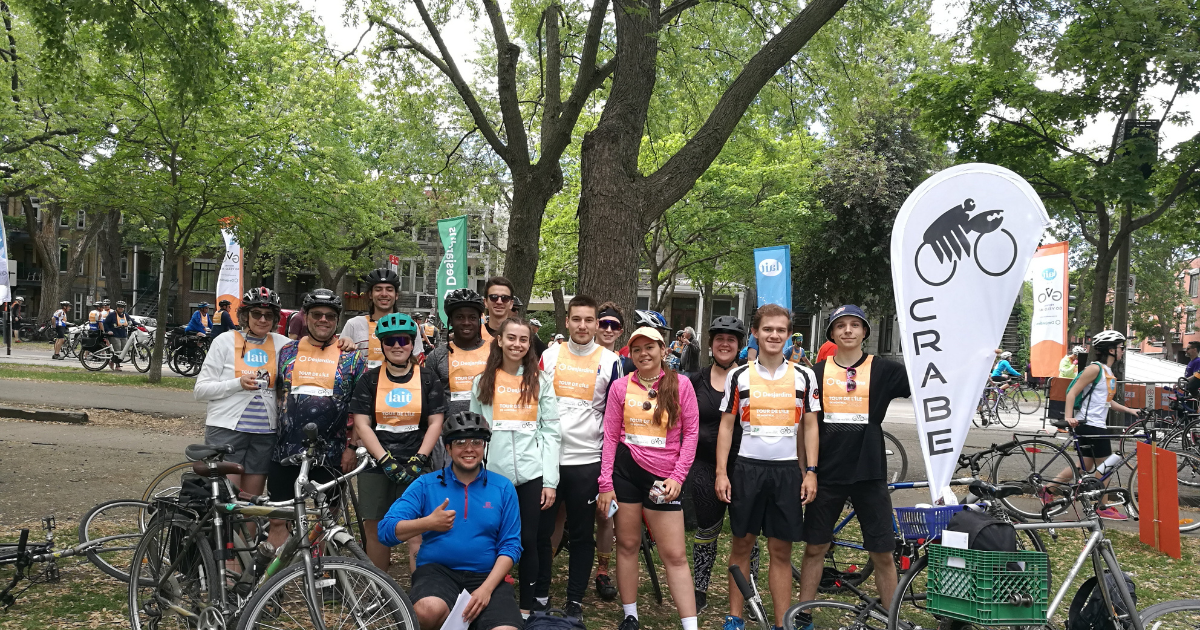
Fin de session et recherche de stage
16 avril 2024
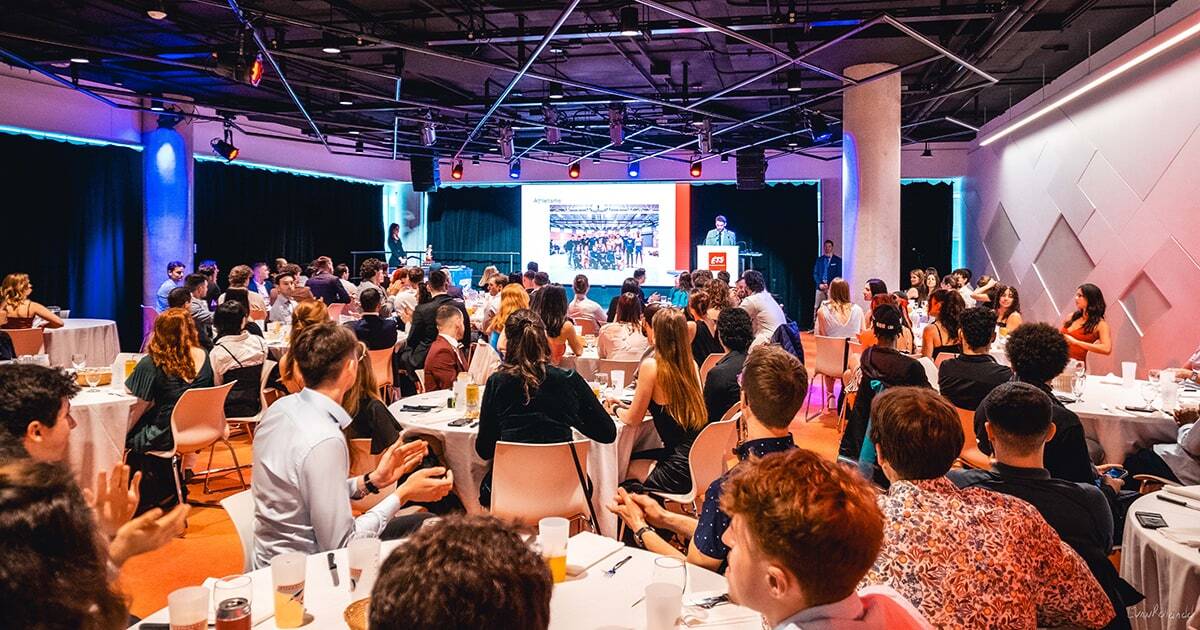
Gala d’excellence : recherche et enseignement 2024
11 avril 2024
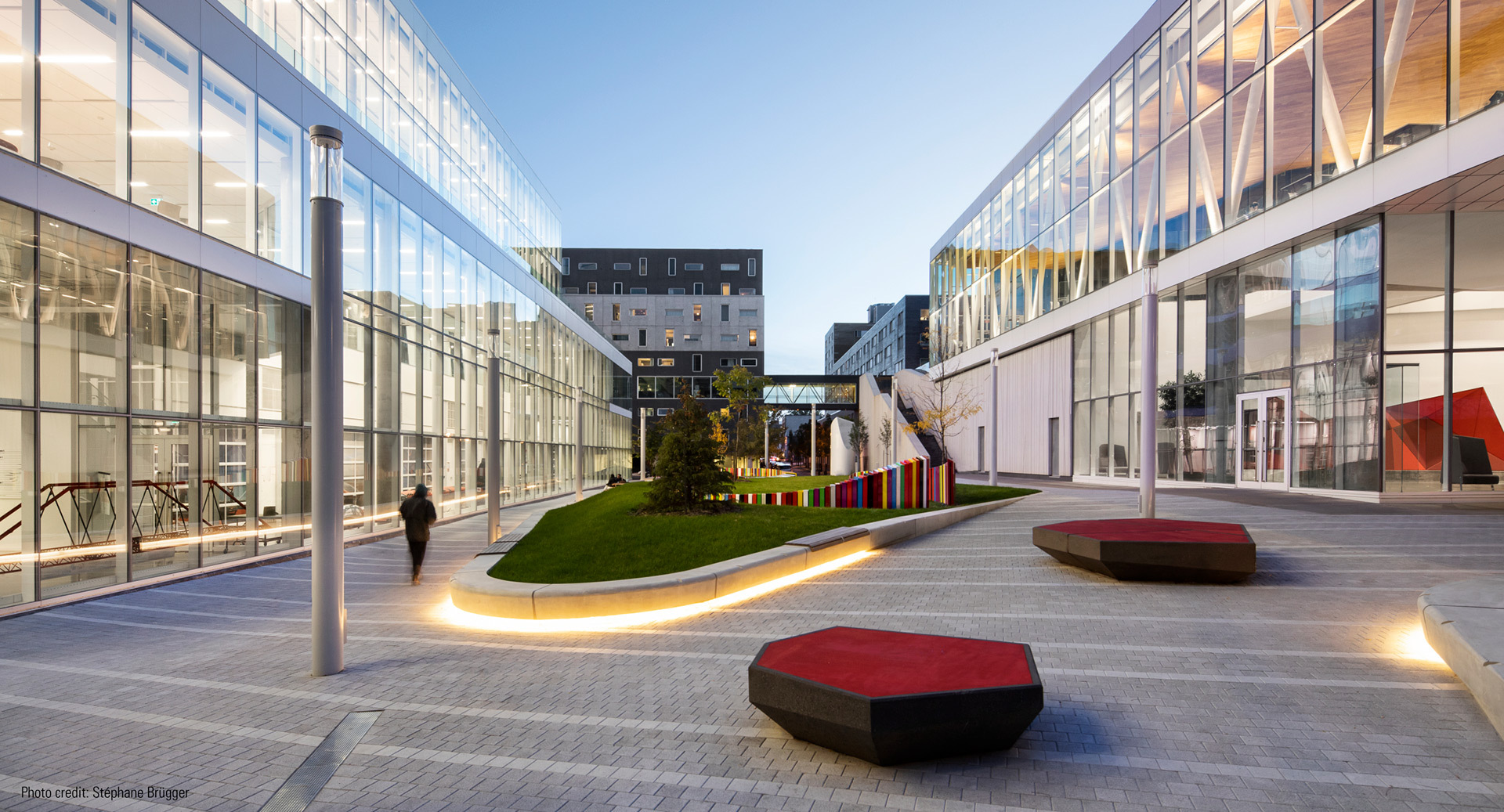
L’ÉTS accueillera la 92ᵉ édition du Congrès de l’Acfas avec la collaboration de l’Université Concordia
11 avril 2024
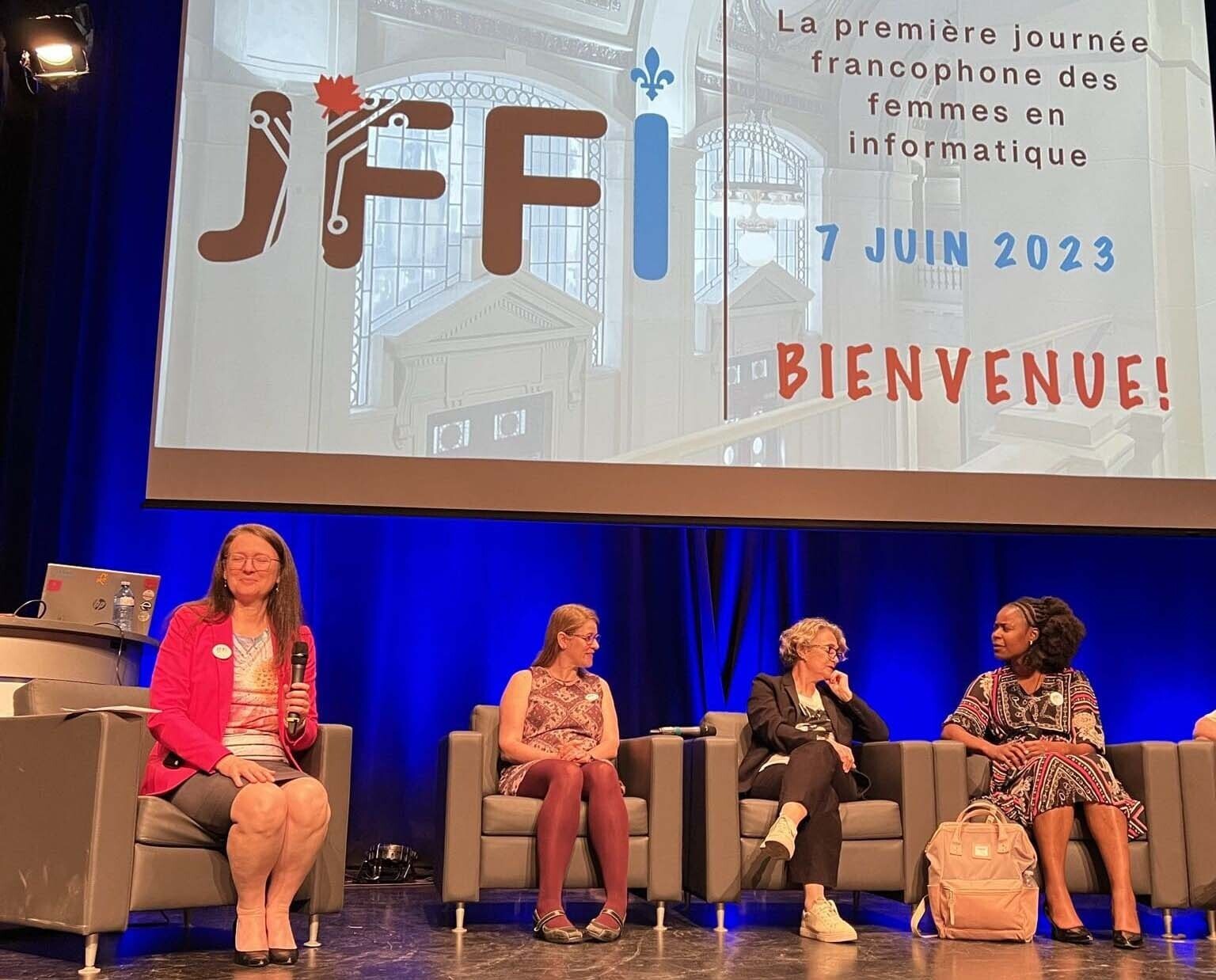
Journée francophone des femmes en informatique
1
Vous êtes actuellement sur cette page
2
Aller à la page : 2
3
Aller à la page : 3
...
51
Aller à la page : 51
Aller à la page suivante
Explorez votre avenir
universitaire